Gene Expression Analysis Platform Evaluation for FFPE Specimen Material-Based Studies
Email Principal Investigator

Mateusz Koptyra
CBTN Specimen
CBTN Participants
Backer
CHOP Center for Data-Driven Discovery in Biomedicine
About this
Project
Understanding pediatric brain cancers and developing new treatments requires constant innovation. In this project, researchers have developed a new technology for the analysis of tumor RNA, particularly mRNA, the strands of RNA that carry messages through a cell. This new method is called the HTG EdgeSeq technology pipeline for mRNA and/or miRNA profiling. HTG technology is able to detect the expression of 2568 genes from different sample types. HTG EdgeSeq is usable for small clinical specimens and according to the manufacturer it requires only a small amount of sliced tissue (formalin-fixed paraffin-embedded or FFPE) to obtain quality mRNA profiling. Researchers will analyze FFPE material from primitive neuroectodermal tumors, high grade gliomas, and medulloblastoma samples using HTG Oncology Biomarker Panel. To assess the HTG system’s ability to profile mRNA from FFPE, researchers are looking to analyze samples that have already undergone RNA sequencing. The Children’s Brain Tumor Network provides a unique resource of FFPE samples where RNAsequencing, an analysis of the RNA in a sample, was already completed. By comparing CBTN data with results from this work, researchers can validate their new RNA analysis platform appropriately. Potential implementation of such technology in the center may allow to obtain the quality data from highly degraded, and of very limited availability material specimens.
Ask The
Scientists
What are the goals of this project?
Researchers will use CBTN samples to assess whether the HTG system can properly profile mRNA from FFPE.
What is the impact of this project?
This technology could allow for the extraction of high quality data from even highly degraded or limited specimen samples, advancing the field of cancer biology.
Why is the CBTN request important to this project?
The Children’s Brain Tumor Network already conducted RNA sequencing on samples provided to this project. Use of such samples will allow researchers to verify the usefulness of HTG technology for this purpose.
Specimen Data
The Children's Brain Tumor Network contributed to this project by providing slides for the generation of miRNAseq data.
Meet The
Team
related
Histologies
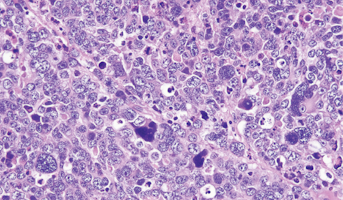
Medulloblastoma
Medulloblastomas comprises the vast majority of pediatric embryonal tumors and by definition arise in the posterior fossa, where they constitute approximately 40% of all posterior fossa tumors. Other forms of embryonal tumors each make up 2% or less of all childhood brain tumors.The clinical feature
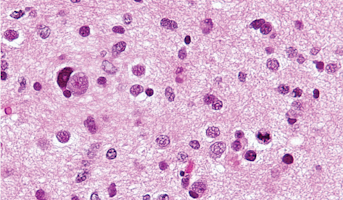
High-Grade Glioma
High-grade Gliomas (HGG) or astrocytomas in children nearly always result in a dismal prognosis. Although novel therapeutic approaches are currently in development, preclinical testing has been limited, due to a lack of pediatric-specific HGG preclinical models. These models are needed to help test




