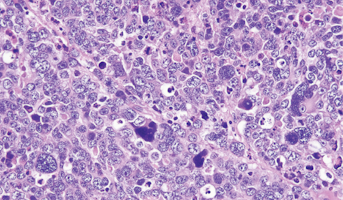
Integrative Functional Genomics of Recurrent Childhood Medulloblastoma
Email Principal Investigator

Javad Nazarian
CBTN Data
CBTN Specimen
CBTN Participants
About this
Project
Medulloblastoma (MB), the most common malignant brain tumor in children, often confers a poor prognosis, especially for patients who have cancer relapse. Tumor relapse is among the most powerful predictors of prognosis despite therapeutic intervention and clinical trial enrollment in patients with MB. Little is known about the molecular genetics of MB relapse and although MB subgroup status remains stable at relapse, the degree that the genetics is altered between these disease states is unclear. Other entities have proposed that MB relapses spawn from rare, treatment-resistant subclones present in the primary tumor, but this has yet to be substantiated. Since MB clinical trials will begin to evaluate targeted therapies in patients with relapsed MB, understanding how MB relapses are related to the primary tumors from which they arise is integral to the success of therapies. Systematic, genome-wide studies designed to effectively investigate the molecular relationship between diagnostic and relapsed MB in a sizable cohort are needed. This project seeks to do just that by utilizing tumor samples and access to the Pediatric Brain Tumor Atlas, both provided by the Children’s Brain Tumor Network.
Ask The
Scientists
What are the goals of this project?
Researchers on this project aim to understand how MB relapses are related to the primary tumors from which they arise and the mechanisms for how tumors evolve and metastasize.
What is the impact of this project?
Samples and data on relapsed MBs are rare, so access to the Pediatric Brain Tumor Atlas and samples from the Children’s Brain Tumor Network are invaluable.
Specimen Data
The Children's Brain Tumor Network contributed to this project by providing tumor DNA and access to the Pediatric Brain Tumor Atlas
Meet The
Team
Madhuri Kambhampati, technician

Washington, DC, USA

Washington, DC, USA

262 Danny Thomas Place, Memphis, TN 38105
Institutions

Primary
Children’s National Hospital
Joined onEach year, the Brain Tumor Institute at Children’s National evaluates more than 100 new patients with brain tumors, and is recognized as a world leader in childhood brain tumor care and research. Children’s National has pioneered novel pediatric brain tumor therapies, including new molecularly-targe


St. Jude Children's Research Hospital
St. Jude is leading the way the world understands, treats and defeats childhood cancer and other life-threatening diseases. The mission of St. Jude Children’s Research Hospital is to advance cures, and means of prevention, for pediatric catastrophic diseases through research and treatment. Consisten