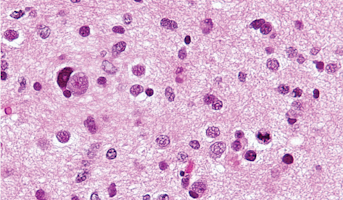
Targeting Replicative Stress in Pediatric Brain Tumors with ALT
Email Principal Investigator

Kristina A. Cole
CBTN Specimen
CBTN Participants
CBTN Pre-clinical Models
Backer
Internal funding
About this
Project
Wee1 is a kinase molecule that plays a big role in the regulation of mitosis, the process of cell division. Many cancers are associated with the overexpression of Wee1, making it a possible target for therapies. AZD-1775, an inhibitor molecule, is under investigation as a radiation sensitizer for children with newly diagnosed DIPG due to Wee1 overexpression. Radiation sensitizers are substances whose use can enhance the effectiveness of radiation therapy. Researchers hypothesize that ATR, CHK1 and Wee1 inhibitors will have single agent activity and radiation sensitization properties in subsets of pediatric supratentorial PNETs and HGG. The goals of this project are to test ATR, CHK1 and Wee1 inhibition (alone and in combination with radiation therapy) in pediatric brain tumor cell lines with and without ALT. If pHGGs and sPNET with ALT are inhibited with ATRi, CHK1i, and this augments radiation sensitivity, it would provide the evidence needed to bring these drugs to the next phase of clinical trial. Since ALT is also found in adult lower grade glioma, pancreatic neuroendocrine tumor, neuroblastoma, and subtypes of sarcoma, this work could also have a broader application across the field of cancer therapy. Researchers will use specimens provided by the Children’s Brain Tumor Network to create the models needed to complete this research.
Ask The
Scientists
What are the goals of this project?
Researchers will test ATR, CHK1, and Wee1 inhibition as a possible therapeutic option for patients with pediatric PNETS and HGGs.
What is the impact of this project?
This work could have an immediate impact on the advancement of pediatric brain cancer treatments, but it is also more broadly applicable across many types of cancer as well.
Why is the CBTN request important to this project?
The high quality specimens provided by the Children’s Brain Tumor Network allow researchers to develop the models needed to complete this work.
Specimen Data
The Children's Brain Tumor Network contributed to this project by providing tumor and germline DNA, tissue for cell line generation and cell lines.
Meet The