Jo Lynne Rokita

About
Supervisory Bioinformatics Scientist
Children's Hospital of Philadelphia
Jo Lynne’s broad research interests lie in identifying and validating novel coding, noncoding, epigenetic, and/or transcriptional mechanisms contributing to oncogenesis, tumor evolution, and progression/relapse of pediatric high-grade diffuse astrocytic tumors. She combines molecular, genomics, and machine learning approaches to discover and validate novel actionable oncogenic targets and/or pathways with the overall goal of informing current therapeutic regimens leading to future clinical trials.

Children’s Hospital of Philadelphia
scientific
Projects

Data
Ongoing
Telomere Maintenance Across Multiple Brain Tumors
A greater understanding of the abundance of telomeres and their relationship to the growth of high grade gliomas (HGG) could lead to new therapies. Using the Pediatric Brain Tumor Atlas, this study seeks to build this knowledge for use by researchers around the world.
HGG

Kristina A. Cole

Data
Ongoing
Molecular Mechanisms and Functional Impact of Aberrant Splicing in Diffuse Midline Gliomas
A comprehensive approach is needed to advance the understanding and clinical care options for patients with diffuse midline gliomas (DMGs). Using DMG samples provided by the Children’s Brain Tumor Network, researchers seek to identify avenues for new therapies.
DIPG

Ammar Naqvi
research
Interests
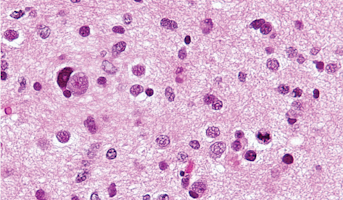
High-Grade Glioma
High-grade Gliomas (HGG) or astrocytomas in children nearly always result in a dismal prognosis. Although novel therapeutic approaches are currently in development, preclinical testing has been limited, due to a lack of pediatric-specific HGG preclinical models. These models are needed to help test
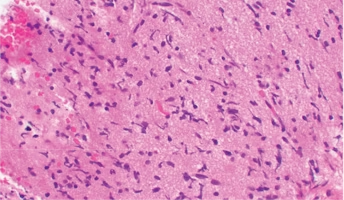
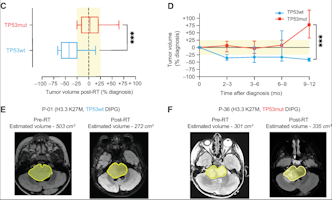
Diffuse Intrinsic Pontine Glioma
A presumptive diagnosis of DIPG based on classic imaging features, in the absence of a histologic diagnosis, has been routinely employed. Increasingly however, histologic confirmation is obtained for both entry into research studies and molecular characterization of the tumor.[5] New approaches with